Below are selected publications. You will find a full record here.
2024
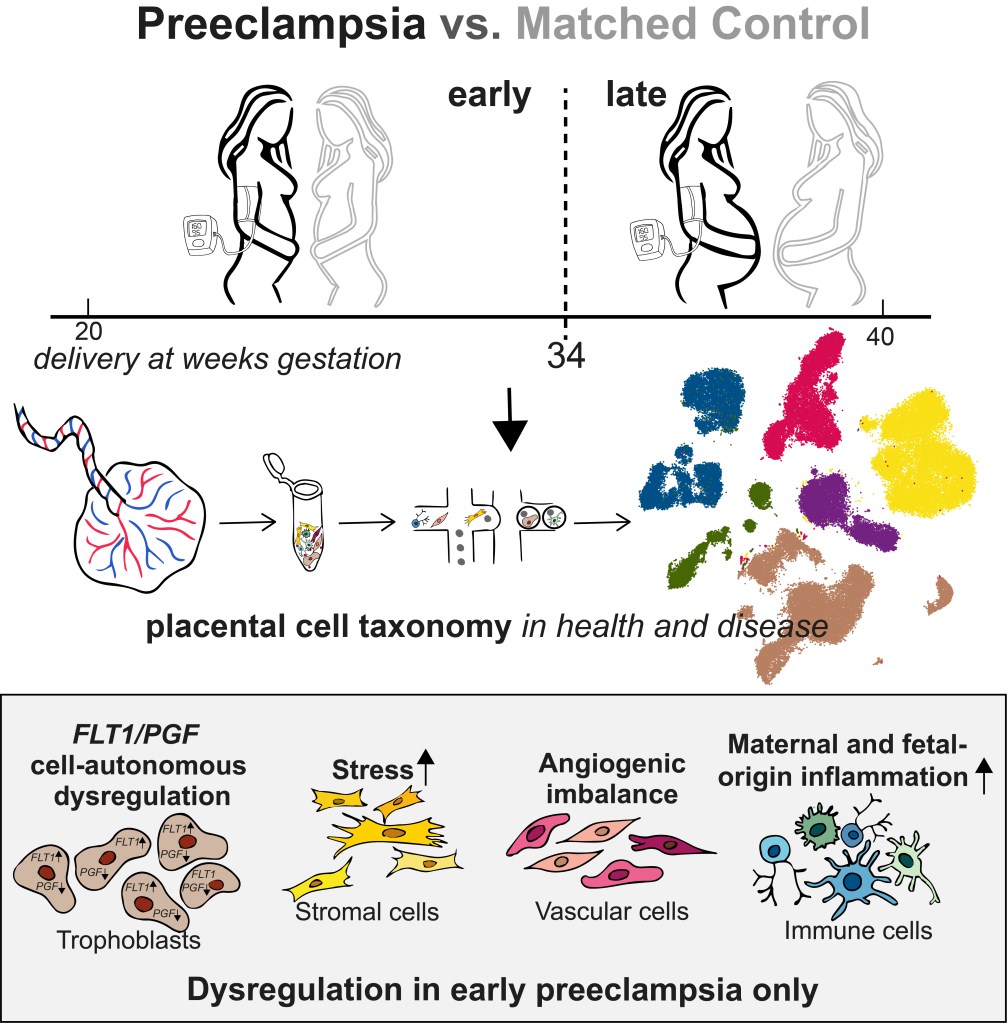
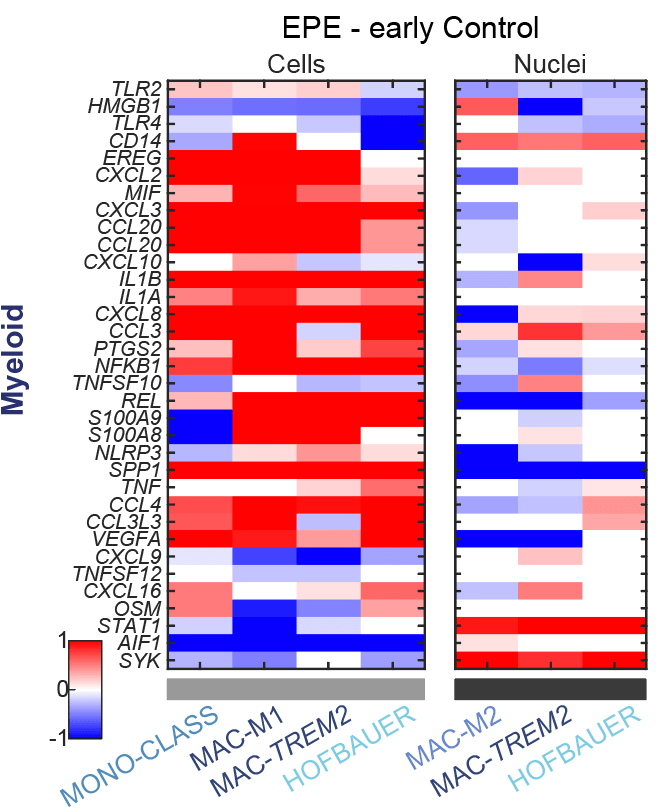
Single-nuclei RNA-sequencing fails to detect molecular dysregulation in the preeclamptic placenta. Admati I, Skarbianskis N, Hochgerner H, Ophir O, Yagel S, Solt I, Zeisel A.
Published in Placenta.
Resources: Explore single-cell and single-nuclei expression across placental cell types. Our code is on GitHub.
2023
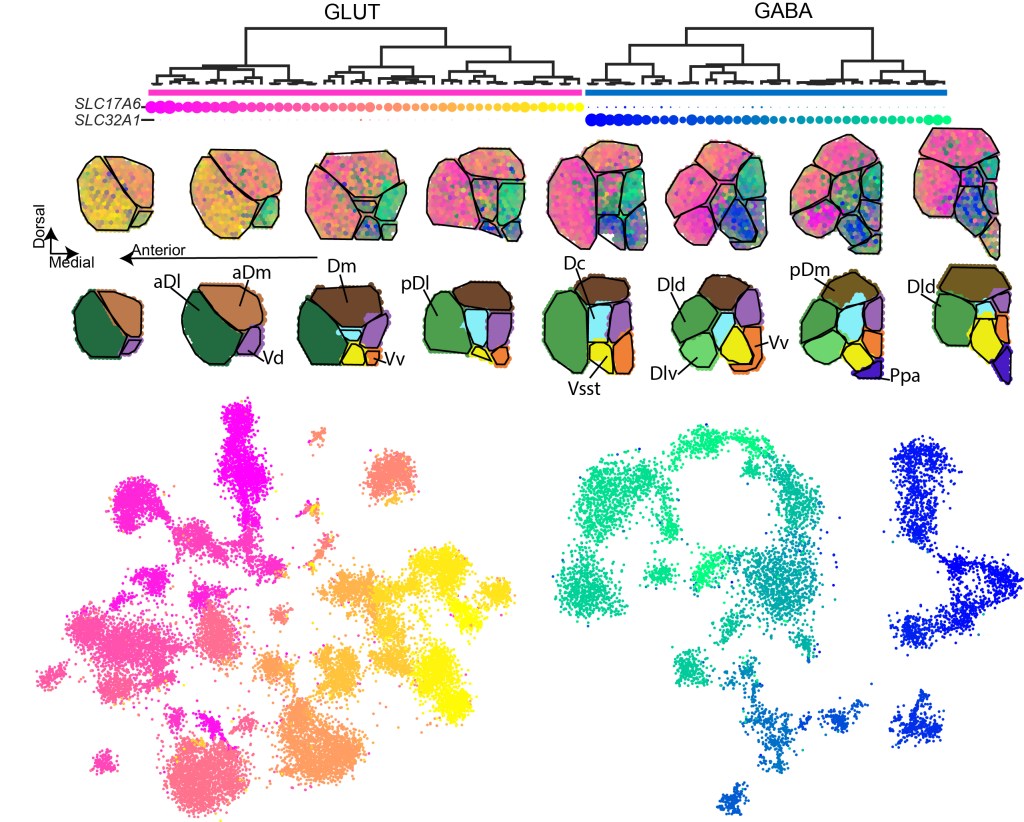
A telencephalon cell type atlas for goldfish reveals diversity in the evolution of spatial structure and cell types. Tibi M, Biton S, Hochgerner H, Lin Z, Givon S, Ophir O, Shay T, Mueller T, Segev R, Zeisel A.
Published in Science Advances. | PDF
Resources: Explore single-cell and spatial gene expression and cell types. And our code is on GitHub.
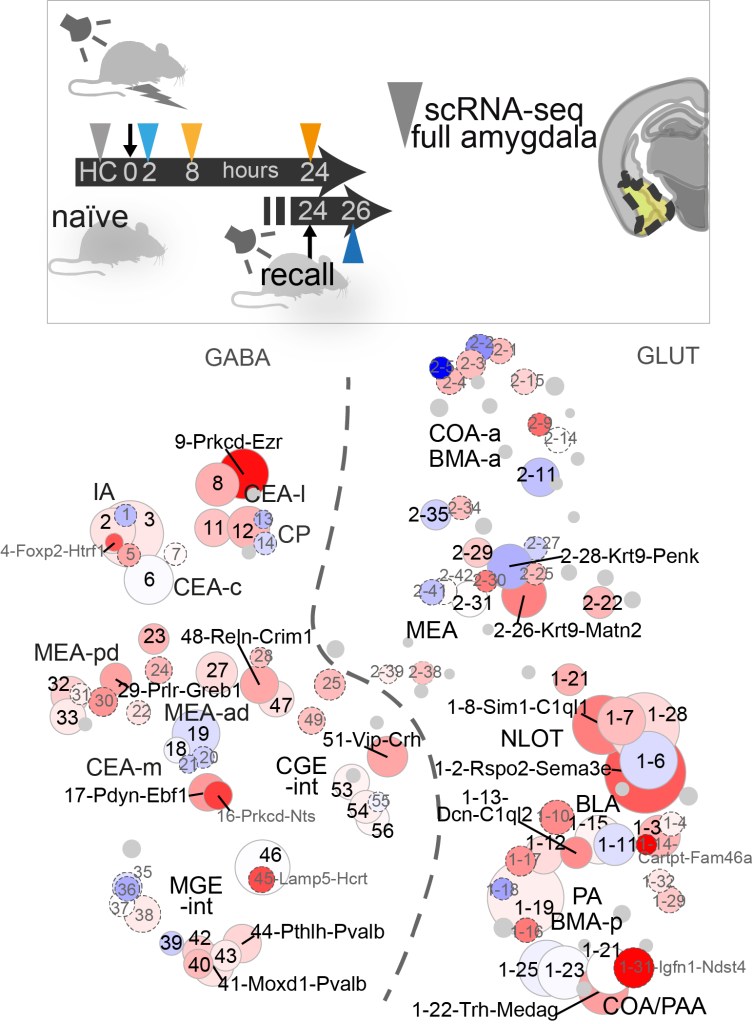
Neuronal types in the mouse amygdala and their transcriptional response to fear conditioning. Hochgerner H, Singh S, Tibi M, Lin Z, Skarbianskis N, Admati I, Ophir O, Reinhardt N, Netser S, Wagner S & Zeisel A
Published in Nature Neuroscience (open access).
Resources | Single-cell gene expression: Cell type explorer and UMI count table. Our code is on GitHub.
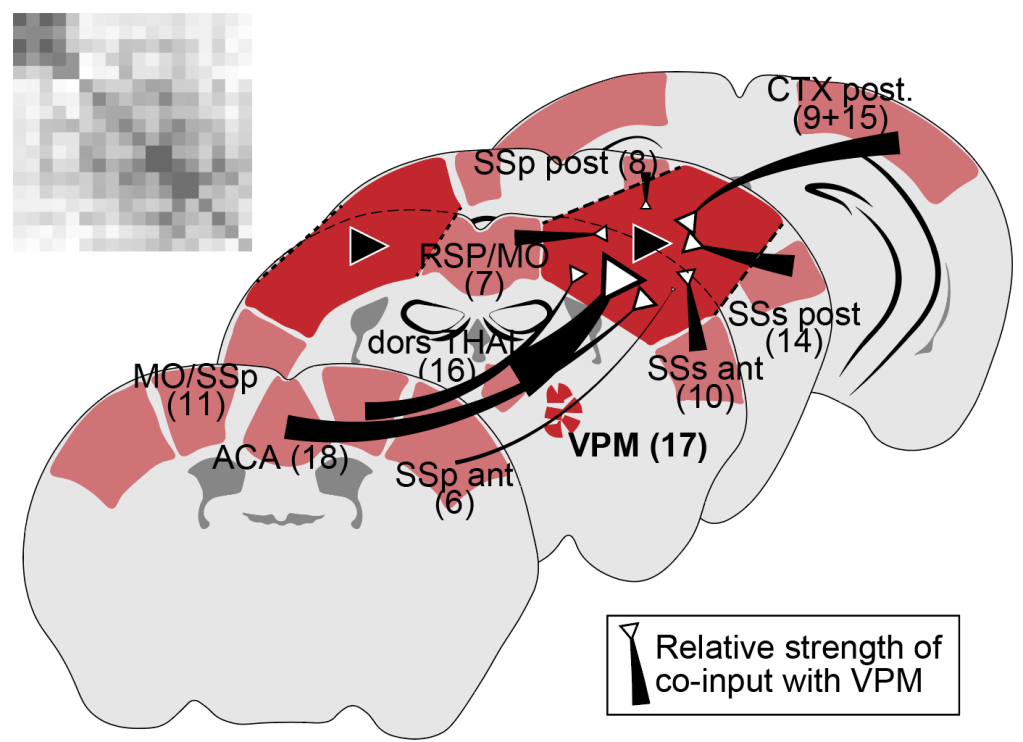
Sexual dimorphism in synaptic inputs to the mouse amygdala and orbital cortex. Aloni E, Tibi M, Hochgerner H, Zeisel A
Published in Frontiers in Neuroscience.

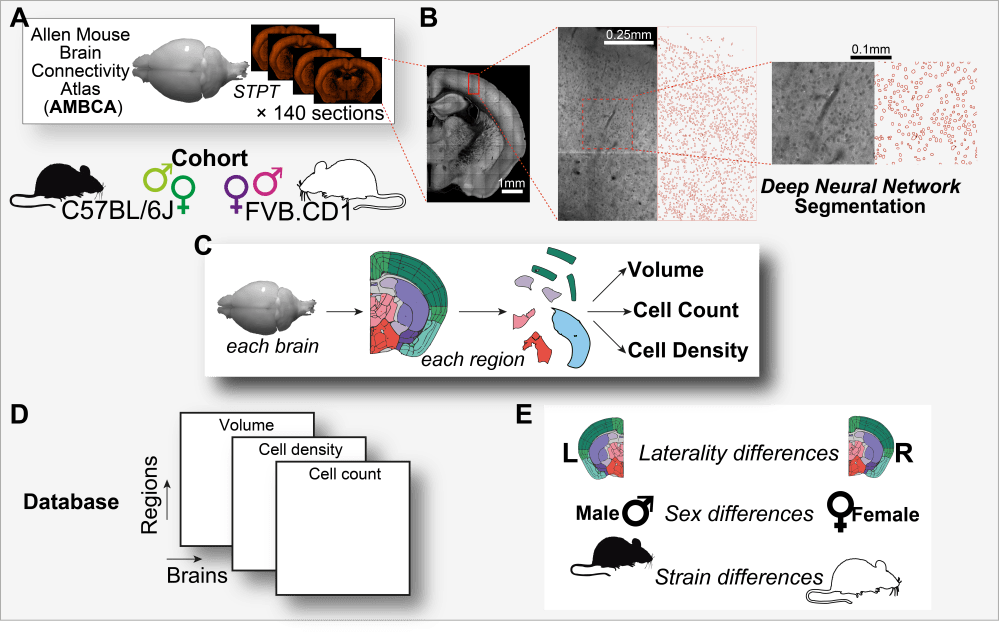
Sex, strain, and lateral differences in brain cytoarchitecture across a large mouse population. Elkind D, Hochgerner H, Aloni E, Shental N, Zeisel A
Published in eLife.
Resources: Brain Data Explorer and GitHub
2022
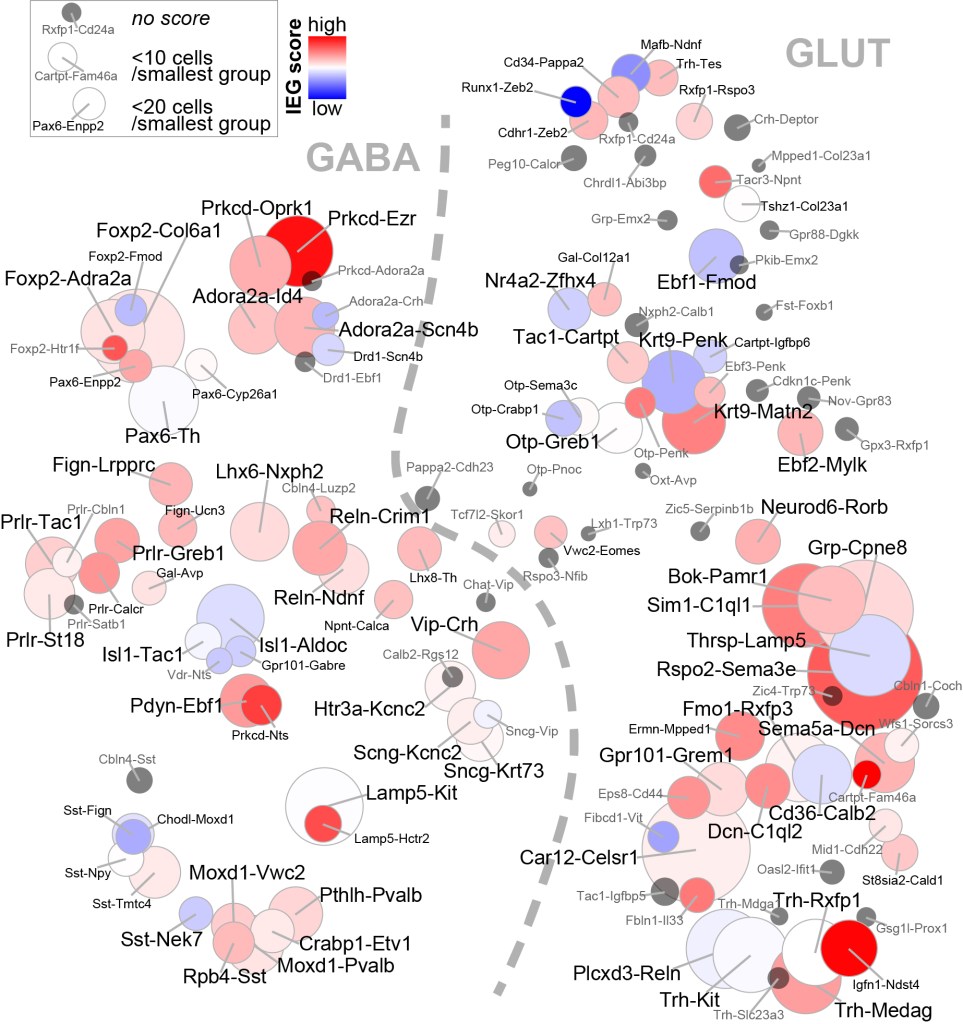
PREPRINT. Cell types in the mouse amygdala and their transcriptional response to fear conditioning. Hochgerner et al.
Preprint available from the bioRxiv | PDF
The publication is available open access on Nature Neuroscience, and you can explore gene expression across the cell atlas here.
…
Hello relocation, establishing a lab, Covid-19 pandemic, and parenthood…
…
2018
Molecular Architecture of the Mouse Nervous System. Published in Cell. Zeisel A, Hochgerner H, Lönnerberg P, Johnsson A, Memic F, van der Zwan J, Häring M, Braun E, Borm LE, La Manno G, Codeluppi S, Furlan A, Lee K, Skene N, Harris KD, Hjerling-Leffler J, Arenas E, Ernfors P, Marklund U, Linnarsson S
PDF
Resources: Adolescent Mousebrain Atlas, at mousebrain.org
RNA velocity of single cells. Published in Nature. La Manno G, Soldatov R, Zeisel A, Braun E, Hochgerner H, Petukhov V, Lidschreiber K, Kastriti ME, Lönnerberg P, Furlan A, Fan J, Borm LE, Liu Z, van Bruggen D, Guo J, He X, Barker R, Sundström E, Castelo-Branco G, Cramer P, Adameyko I, Linnarsson S, Kharchenko PV.
Neuronal atlas of the dorsal horn defines its architecture and links sensory input to transcriptional cell types. Published in Nature Neuroscience. Häring M*, Zeisel A*, Hochgerner H*, Rinwa P, Jakobsson JET, Lönnerberg P, La Manno G, Sharma N, Borgius L, Kiehn O, Lagerström MC, Linnarsson S, Ernfors P.
PDF
Resources: Data+visualization at linnarssonlab.org
Conserved properties of dentate gyrus neurogenesis across postnatal development revealed by single-cell RNA sequencing. Published in Nature Neuroscience. Hochgerner H*, Zeisel A*, Lönnerberg P, Linnarsson S.
PDF
Resources: Data+visualization at linnarssonlab.org
2017
STRT-seq-2i: dual-index 5′ single cell and nucleus RNA-seq on an addressable microwell array. Published in Scientific Reports. Hochgerner H, Lönnerberg P, Hodge R, Mikes J, Heskol A, Hubschle H, Lin P, Picelli S, La Manno G, Ratz M, Dunne J, Husain S, Lein E, Srinivasan M, Zeisel A, Linnarsson S.
Molecular interrogation of hypothalamic organization reveals distinct dopamine neuronal subtypes. Published in Nature Neuroscience. Romanov RA*, Zeisel A*, Bakker J, Girach F, Hellysaz A, Tomer R, Alpár A, Mulder J, Clotman F, Keimpema E, Hsueh B, Crow AK, Martens H, Schwindling C, Calvigioni D, Bains JS, Máté Z, Szabó G, Yanagawa Y, Zhang MD, Rendeiro A, Farlik M, Uhlén M, Wulff P, Bock C, Broberger C, Deisseroth K, Hökfelt T, Linnarsson S, Horvath TL, Harkany T.
2016
Single-Cell Transcriptomics Reveals that Differentiation and Spatial Signatures Shape Epidermal and Hair Follicle Heterogeneity. Published in Cell Systems. Joost S, Zeisel A, Jacob T, Sun X, La Manno G, Lönnerberg P, Linnarsson S*, Kasper M*
Oligodendrocyte heterogeneity in the mouse juvenile and adult central nervous system. Published in Science. Marques S, Zeisel A, Codeluppi S, van Bruggen D, Mendanha Falcão A, Xiao L, Li H, Häring M, Hochgerner H, Romanov RA, Gyllborg D, Muñoz-Manchado AB, La Manno G, Lönnerberg P, Floriddia EM, Rezayee F, Ernfors P, Arenas E, Hjerling-Leffler J, Harkany T, Richardson WD, Linnarsson S*, Castelo-Branco G*.
Integration of electrophysiological recordings with single-cell RNA-seq data identifies neuronal subtypes. Published in Nature Biotechnology. Fuzik J*, Zeisel A*, Máté Z, Calvigioni D, Yanagawa Y, Szabó G, Linnarsson S*, Harkany T*.
2015
Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Published in Science. Zeisel A*, Muñoz-Manchado AB*, Codeluppi S, Lönnerberg P, La Manno G, Juréus A, Marques S, Munguba H, He L, Betsholtz C, Rolny C, Castelo-Branco G, Hjerling-Leffler J, Linnarsson S.
2013
Quantitative single-cell RNA-seq with unique molecular identifiers. Published in Nature Methods. Islam S, Zeisel A, Joost S, La Manno G, Zajac P, Kasper M, Lönnerberg P, Linnarsson S.
2011
Coupled pre‐mRNA and mRNA dynamics unveil operational strategies underlying transcriptional responses to stimuli. Published in Molecular Systems Biology. Zeisel A, Köstler WJ, Molotski N, Tsai JM, Krauthgamer R, Jacob Hirsch J, Rechavi G, Soen Y, Jung S, Yarden Y, Domany E
FDR control with adaptive procedures and FDR monotonicity. Published in The Annals of Applied Statistics. Zeisel A, Zuk O, Domany E
2010
BMP-mediated inhibition of FGF signaling promotes cardiomyocyte differentiation of anterior heart field progenitors. Published in Development. Tirosh-Finkel L, Zeisel A, Brodt-Ivenshitz M, Shamai A, Yao Z, Seger R, Domany E, Tzahor E.